WHS Oral Abstracts
(WHS-L3.01) Resolution of Bioburden and Wound Closure in a Porcine Full Thickness Wound Model
Thursday, May 16, 2024
10:30 AM - 11:30 AM East Coast USA Time
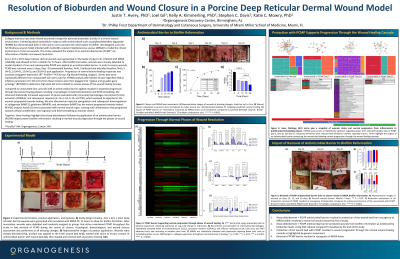
Collagen matrices have been shown to provide a target for aberrant proteolytic activity in a chronic wound environment. Combining these matrices with antimicrobials such as silver or polyhexamethylene biguanide (PHMB) has demonstrated both in vitro and in vivo to prevent the reformation of biofilm. However, a broader understanding of wounds treated with these devices is limited. We designed a porcine full thickness wound model infected with methicillin-resistant Staphylococcus aureus (MRSA) to model the clinical environment of chronic wounds. Evaluation of the impact of a native cross-linked collagen matrix with PHMB (PCMP^) on biofilm reformation, wound closure, and gene expression with STRING and Gene Ontology (GO) term assessment over the course of treatment was performed. 4cm x 4cm x 3mm full thickness wounds were generated on the backs of pigs (n=3), infected with MRSA (USA300), and allowed to form a biofilm for 72 hours. After biofilm formation, wounds were sharply debrided to model standard of care and subsequently PCMP was applied. PCMP was changed every 5 days, at which point wounds were assessed at Days -3 (unwounded baseline, N=4), 0 (infected and debrided baseline, N=6), 5 (N=2), 10 (N=5), 15 (N=5), and 20 (N=3) post-treatment initiation. Gene expression within the wound bed was assessed using RT2 Profiler™ PCR Arrays (Pig Wound Healing, Qiagen). Genes that were statistically different from unwounded skin were used for STRING analysis with Markov Cluster Algorithm (MCL) inflation parameter of 3. GO terms from these clusters were then plugged into “reduce and visualize gene ontology” (REVIGO) to determine high level GO terms related to various phases of the wound healing process. Compared to unwounded skin, two pathways were identified: macrophage recruitment/activation and ECM remodeling. We observed statistically increased expression of genes associated with monocyte/macrophages recruitment (CCL2), activation (CD40LG), and subsequent response (IL-1α, IL-1β, IL-10, and TNF), which trended back towards baseline as the wounds healed and bioburden was eliminated. We also observed statistically greater expression of collagenase (MMP1), gelatinase (MMP9), and stromelysin (MMP3) in the wound that resolved as the wound healed. REVIGO analysis found GO terms associated with normal wound repair, starting with inflammation that progressed towards cellular proliferation and migration and ECM remodeling as wounds closed. Together, these findings highlight that sharp debridement followed by a native cross-linked collagen matrix with PHMB helped prevent biofilm reformation and supported normal wound healing, while the changes in gene expression over the course of closure were characterized. ^PuraPly® AM, Organogenesis, Canton, MA

.jpeg)
