WHS Posters
(WHS-P31) Comparative Analysis Between 16S rRNA NGS vs Conventional Culture associated with the Treatment Outcome of Diabetic Foot Ulcers
Friday, May 17, 2024
7:30 AM - 5:00 PM East Coast USA Time
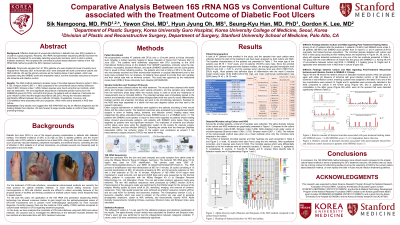
Background: Effective treatment of wound-site infections in diabetic foot ulcer (DFU) patients is crucial for a good prognosis. Recently, 16S rRNA next-generation sequencing (NGS) has been the main focus of research for accurately detecting wound-site microbes, which is vital in optimal antibiotic treatment. We compared the conventional culture-based detection method to the 16S rRNA NGS method to predict the DFU treatment outcomes.
Methods: Wound-site samples from 47 DFU patients who were treated at Korea University Guro Hospital from February 2021 to November 2021 were analyzed with both conventional culture and NGS methods. We set the primary outcome as the healing status of each patient, which was assessed using the SINBAD score and amputation status, and the secondary outcomes as wound-site ischemia and infection control.
Results: The NGS method detected a broader range of microbial species (Shannon index=1.369 ± 0.755, Simpson index=2.987 ± 1.383) compared to the conventional culture method (Shannon index=0.693, Simpson index=1.269). Sixteen species were found using the two methods, which were all anaerobes. The most significant discordance of detected species was found in the SINBAD≥3 group (40.79%), and within that group, the patients with an absence of ischemia but poor infection control had the largest discordance (85.22%). Among the microbes detected significantly different between the two methods, B.fragilis, S.agalactiae, S.aureus, and S.constellatus were associated with poor prognosis, which were mainly detected in NGS than culture.
Conclusion: Early studies now suggest that 16S rRNA NGS may be an effective diagnostic tool for treating diabetic foot infection. We look forward to larger pivotal studies to confirm these initially promising findings.
Methods: Wound-site samples from 47 DFU patients who were treated at Korea University Guro Hospital from February 2021 to November 2021 were analyzed with both conventional culture and NGS methods. We set the primary outcome as the healing status of each patient, which was assessed using the SINBAD score and amputation status, and the secondary outcomes as wound-site ischemia and infection control.
Results: The NGS method detected a broader range of microbial species (Shannon index=1.369 ± 0.755, Simpson index=2.987 ± 1.383) compared to the conventional culture method (Shannon index=0.693, Simpson index=1.269). Sixteen species were found using the two methods, which were all anaerobes. The most significant discordance of detected species was found in the SINBAD≥3 group (40.79%), and within that group, the patients with an absence of ischemia but poor infection control had the largest discordance (85.22%). Among the microbes detected significantly different between the two methods, B.fragilis, S.agalactiae, S.aureus, and S.constellatus were associated with poor prognosis, which were mainly detected in NGS than culture.
Conclusion: Early studies now suggest that 16S rRNA NGS may be an effective diagnostic tool for treating diabetic foot infection. We look forward to larger pivotal studies to confirm these initially promising findings.

.jpeg)
